Medical content
The medical knowledge base (opens in a new tab) is one of the core components of every solution that's developed by Infermedica. It contains thousands of medical concepts - symptoms, conditions, risk factors - that help us describe and precisely identify the most probable issues facing children and adults.
Content development process
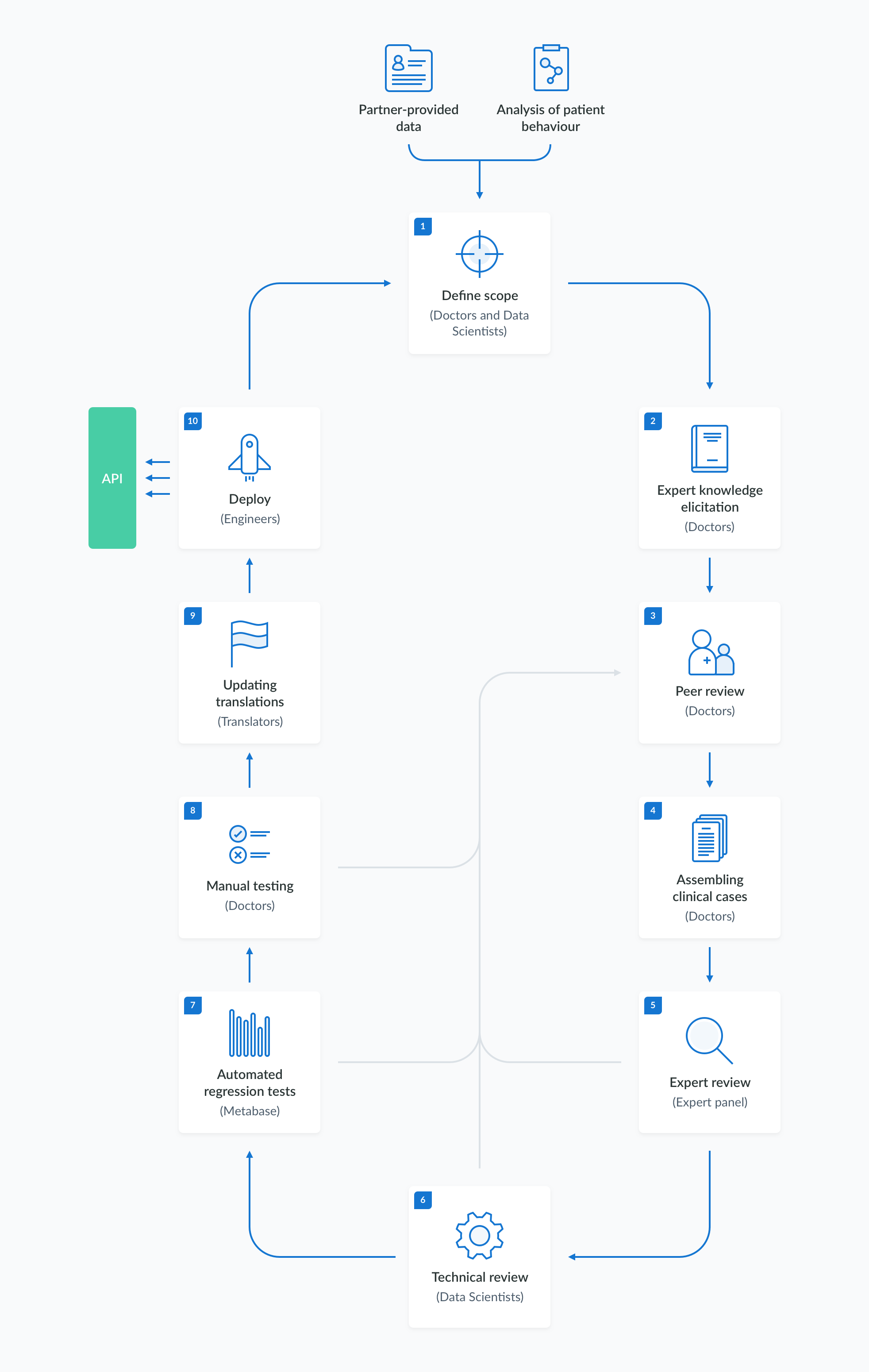
The creation, validation and maintenance of the information in the medical knowledge base follows a rigorous and well-established procedure. The content development process is divided into stages, as outlined in the diagram below. These stages include:
- defining the scope of the desired changes,
- eliciting expert knowledge based on literature and other available sources,
- specifying the test criteria,
- performing manual and automated acceptance tests,
- deploying the verified model to the API.
The process can be efficiently repeated with automated regression testing to ensure the system's stability.
1. Defining scope
Content development begins with defining the scope of the desired changes. With the help of our data scientists, our medical team analyzes statistical information collected by our symptom checkers to discover which areas of our system should be further expanded. This may include adding new pathologies or symptoms, or expanding on the information that's currently available for existing conditions.
The scope definition may also take into account the requirements provided by our customers and partners, who can specify their target audience (e.g. geography, age groups, or range of expected conditions).
2. Expert knowledge elicitation
Once the scope is defined, our medical content editors will collect quality literature regarding the newly introduced topic and use our Metabase to enter evidence-based information in the form of probabilistic characteristics of a given condition’s prevalence, the influence of risk factors, and the sensitivities of symptoms that can be assessed in both categorical and numerical formats. Each piece of introduced medical information contains a reference to the original source, such as a book or article name.
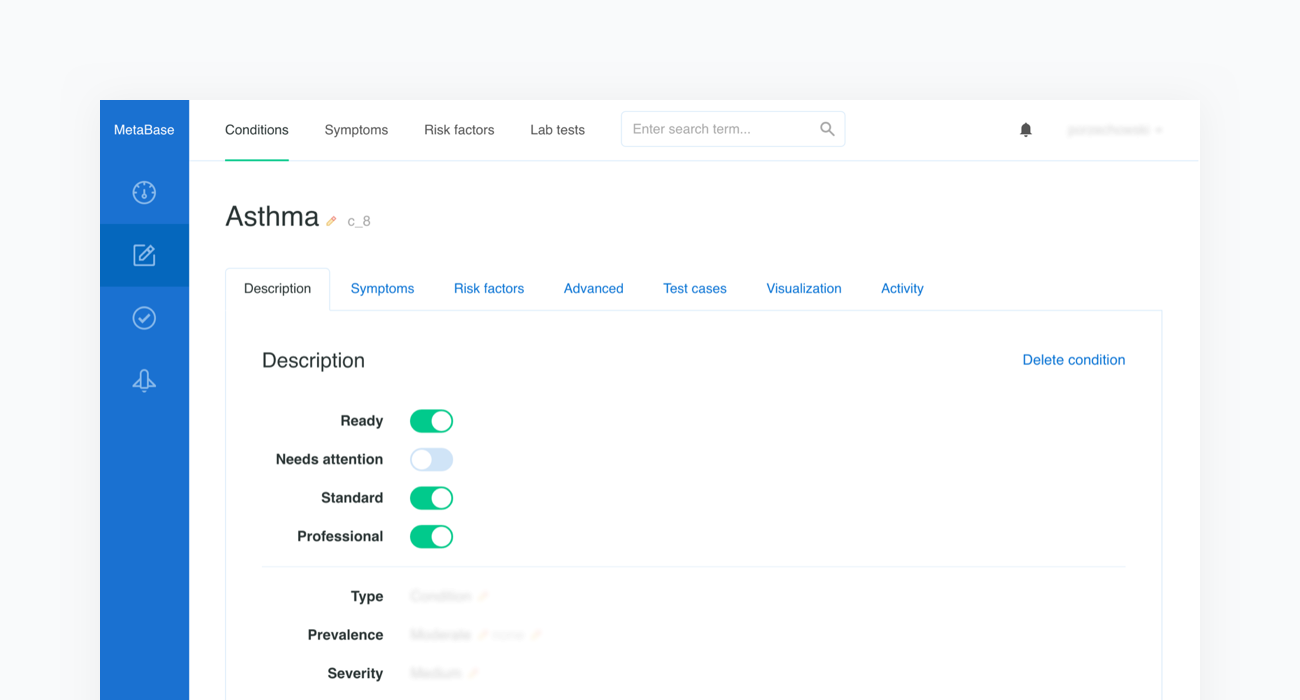
Every change involving the medical content is stored in Metabase's history, which also includes information regarding the time and author. This system makes it possible to track, review, and comment on any change.
3. Peer review
The next step is a peer-review of the new content by another medical content editor. The newly provided data and sources are also validated. Possible improvements are discussed and introduced by editors. Metabase simplifies collaboration between doctors by offering features that enable comments to be exchanged and resolved quickly. Doctors can also notify other editors about their progress by using a system of flags and statuses (e.g. condition in progress, ready for peer-review, requires attention, etc.).
4. Assembling clinical cases
After finalizing the peer-review of the new content, our doctors are responsible for finding and entering literature-based clinical cases that validate the system's performance using the newly provided medical content.
We continually test our medical knowledge base and reasoning algorithm against cases reported in journals, including complex CPC cases published in The New England Journal of Medicine, to ensure adequate performance in every possible clinical presentation.
In order to provide satisfactory performance levels, we've developed a methodology for monitoring and ensuring the quality of the system using real patient clinical cases. In parallel with the development of our medical knowledge base, we've assembled case reports consisting of clinical situation descriptions and a list of expected outcomes (the most probable conditions). We collect patient cases from reputable and well-known sources such as BMJ, NEJM, The American Journal of Medicine, Mayo Clinic Proceedings, BioMed Central, Oxford Journals, and many others, including educational literature (e.g. 100 Cases in Clinical Medicine) as well as United States Medical Licensing Examination Step 2 CK tests. The current results for our clinical test cases, also called acceptance Test Cases (aTC), are presented below.
- conditions accuracy: 96%
- last updated: October 2025
Testing methodology
Construction of a clinical test case begins with finding a source report. In the example below we refer to the case of a “33-Year-Old Woman With Epigastric Pain and Hematemesis”, published in Mayo Clinic Proceedings, Mayo Clin Proc. 2012 Feb; 87(2): 194–197 (opens in a new tab).
The source article is then carefully analyzed by a physician and all the clinical features are extracted before being inserted into a newly created case in our Metabase. The clinical features include the age and sex of the patient as well as any confirmed or excluded findings, such as symptoms and risk factors. Every test case must also contain the resulting condition and the acceptance criteria.
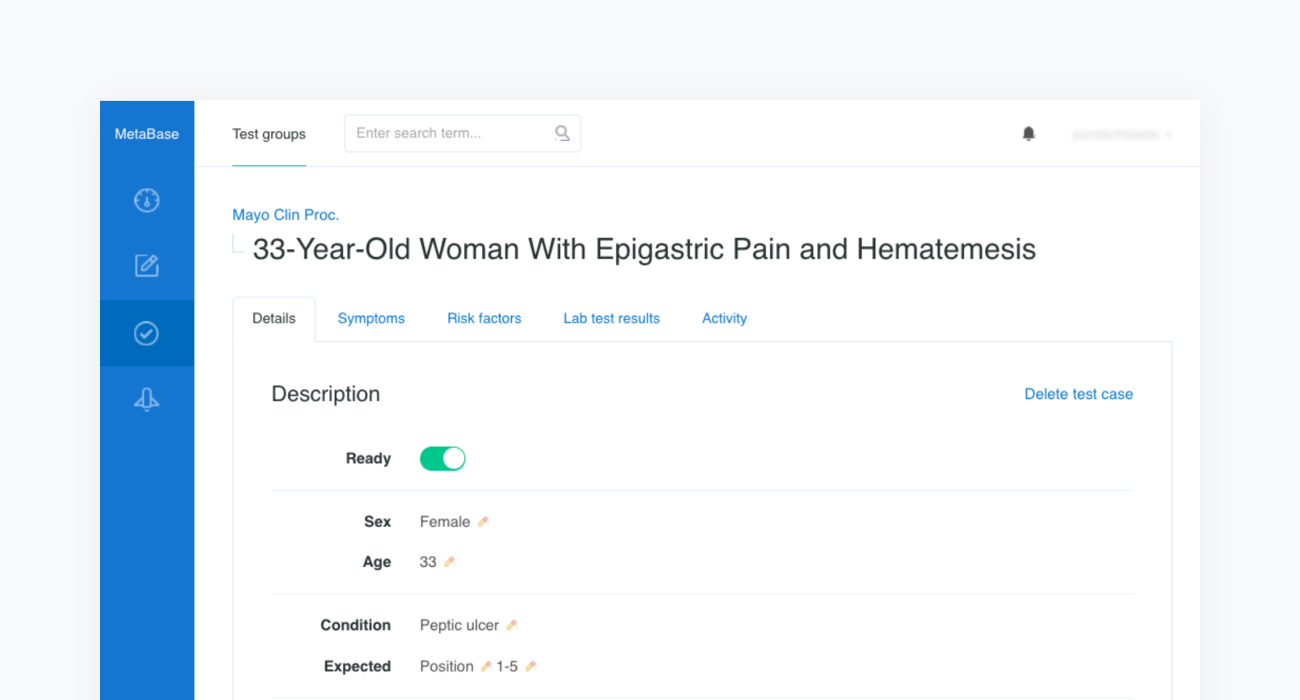
The acceptance criterion applied to each patient case is that the condition is in the top 3 or 5 positions —depending on the complexity of the case— of Infermedica's rankings. Since the goal of the system is to identify a group of likely conditions, rather than suggesting a definitive result, positioning recommendations within such an interval is a reasonable approach that has been used in studies of various CDSS systems.
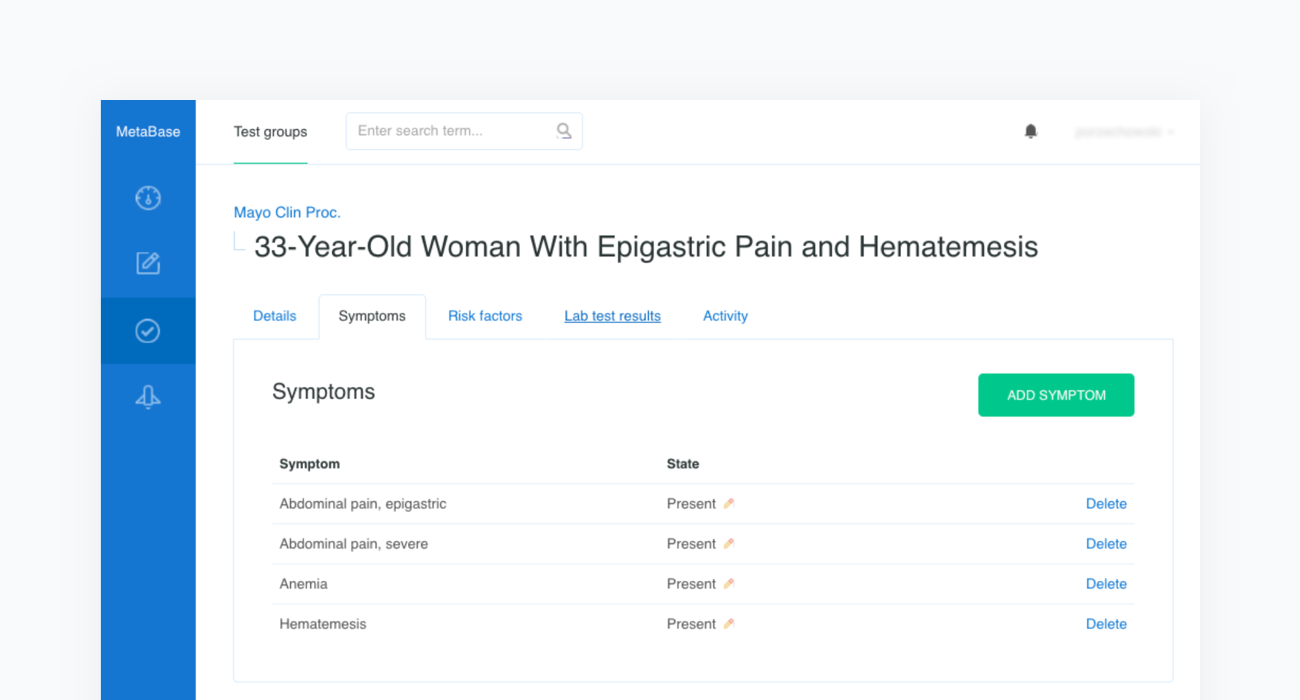
Once all clinical features and test criteria have been provided, Metabase will automatically validate the newly created test case against the inference engine. This procedure is repeated in cycles and serves as a regression testing framework for our system. As shown in the figure below, the case of the “33-Year-Old Woman With Epigastric Pain and Hematemesis” meets the acceptance criteria, as “Peptic ulcer” is listed in the first position of Infermedica's rankings.
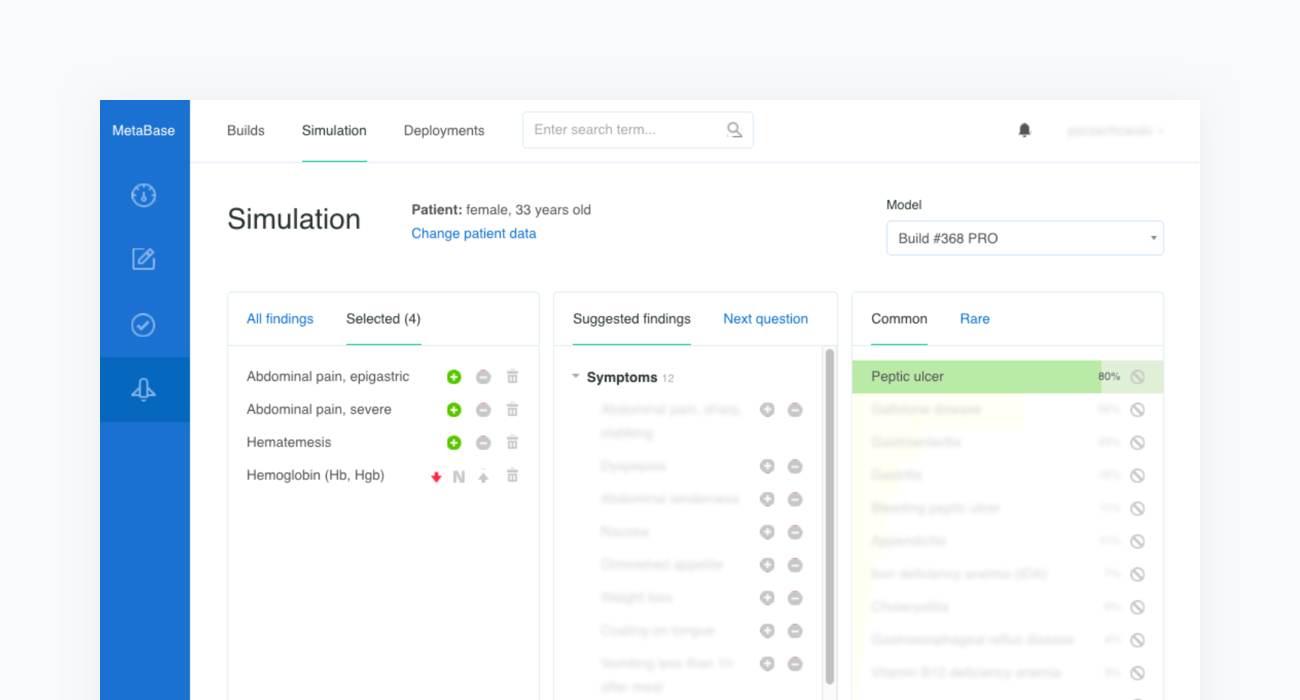
5. Expert review
The newly created content is then verified by a doctor selected from our expert panel who has relevant experience in the given area. Experts can return the process to peer-review in case of any issues found in either the content or clinical cases. When the new content is accepted, the technical review begins.
6. Technical review
The technical review is performed by a data scientist who checks whether the content has been developed according to internal guidelines and identifies any potential issues involving the structuring of the medical content (e.g. checking for duplicated symptoms, verifying the hierarchy of symptoms and numerical parameters introduced by the doctors, etc.). The technical reviewer also works directly with the authors or experts to resolve any issues that arise.
7. Regression testing
At this point the new model is built and all clinical cases are executed. This process is called regression testing. It allows us to test how the newly introduced content influences the performance of the previous model. This guarantees the stability of this complex system and allows us to continuously measure its behavior.
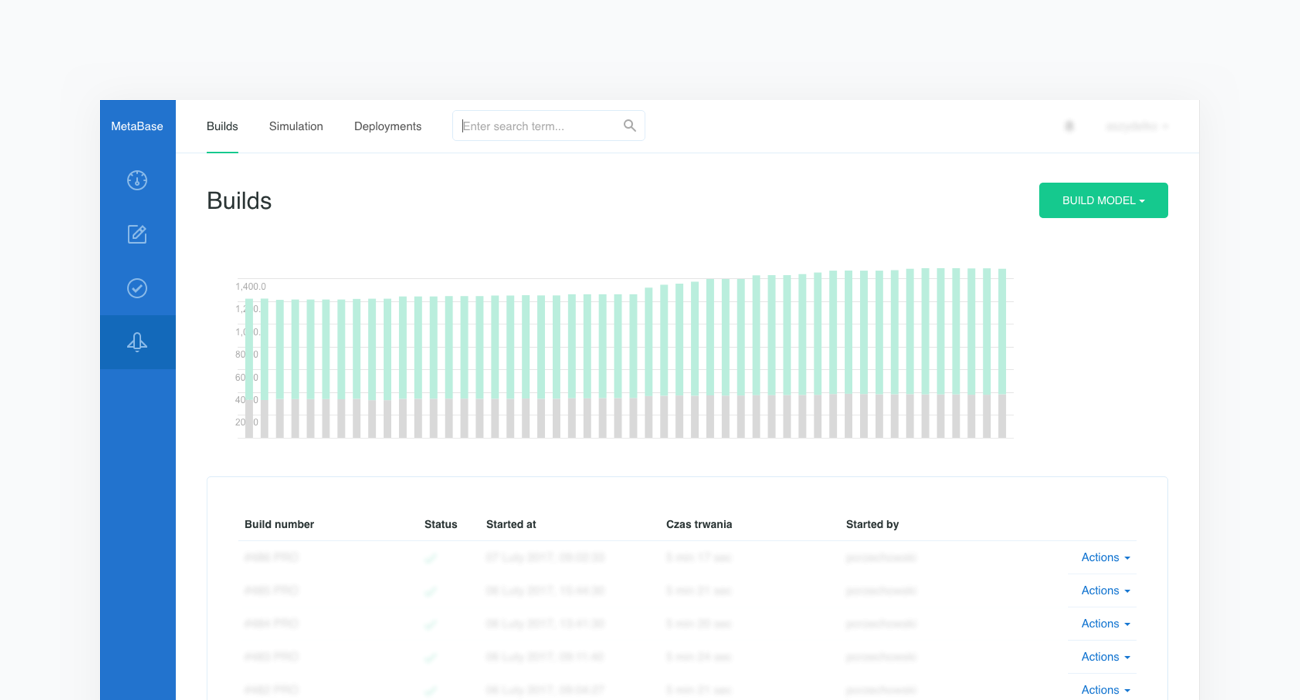
It is important to note that clinical test cases vary in terms of the complexity and rarity of disease representation.
8. Manual testing
Once regression testing is completed and the results have been accepted, our doctors manually test the newly introduced conditions and symptoms. While subjective, this step provides us with an approximation of the real-life experience users have with the Infermedica platform.
9. Updating translations
Metabase content is currently available in Arabic, Chinese - Simplified, Czech, Dutch, English, Estonian, French, German, Greek, Italian, Polish, Portuguese, Portuguese (Brazilian), Romanian, Russian, Slovak, Spanish, Spanish - Latin American, Turkish and Ukrainian. We have extensive experience in preparing localized versions of our system. Translation of the content is provided by a native speaker with medical education and takes about 120-160 hours. With every new content release, we make sure all the language versions are up to date.
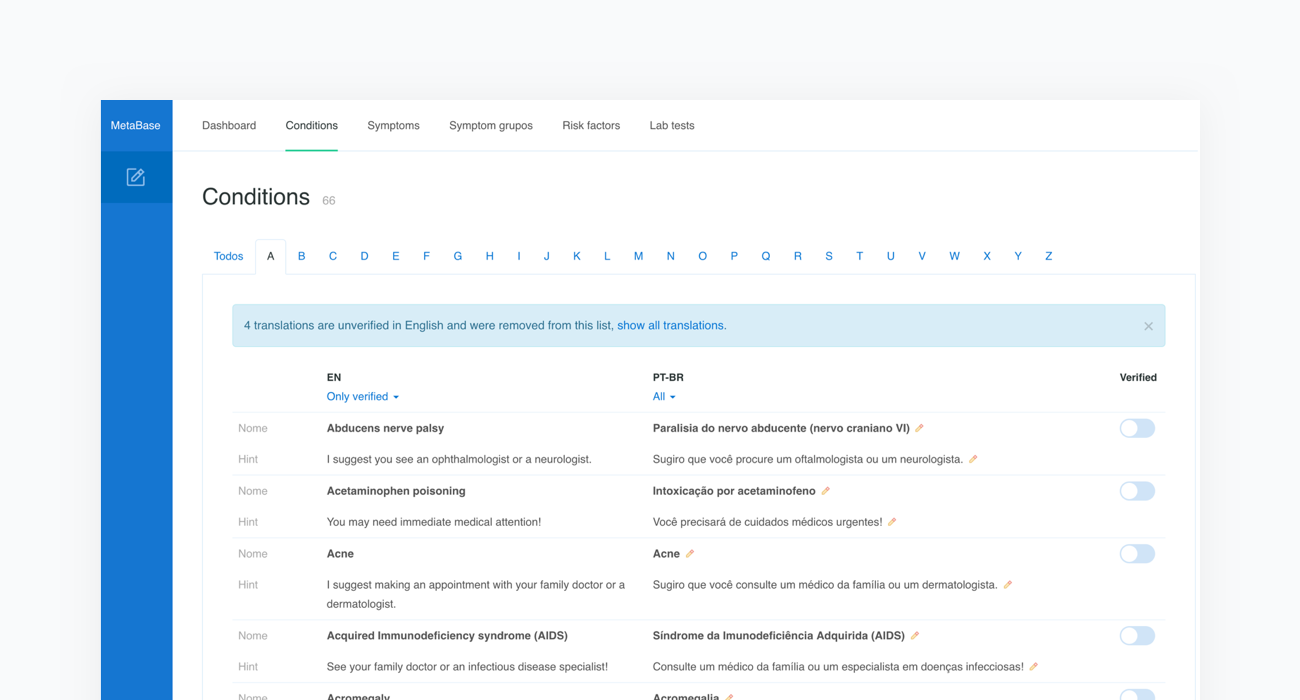
10. Deploy
Lastly, the fully tested models are ready to be deployed to the cloud-based API. Once completed, the new content will be available to all users of Infermedica platform. With every release of a new model, Infermedica provides a fully transparent changelog of the medical content. This allows all of our partners to track modifications as well as the pace at which our medical knowledge base is growing.
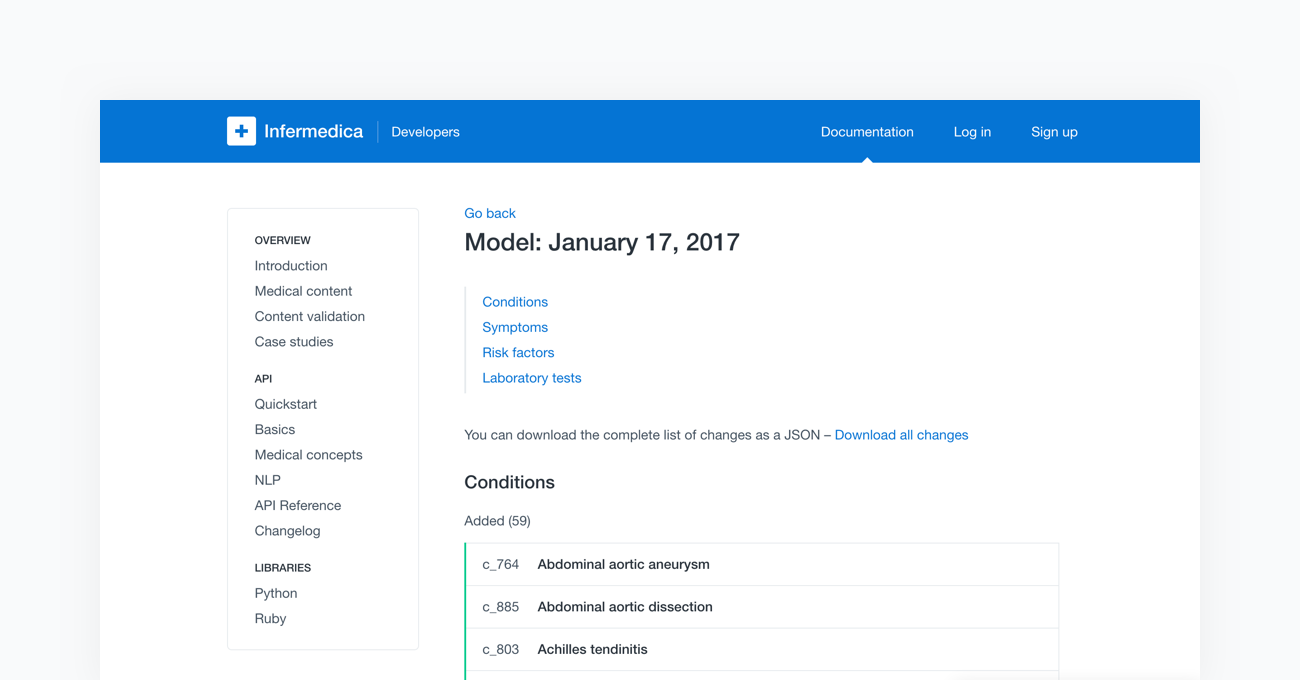
For more information, check out our webpage (opens in a new tab) and blog (opens in a new tab).